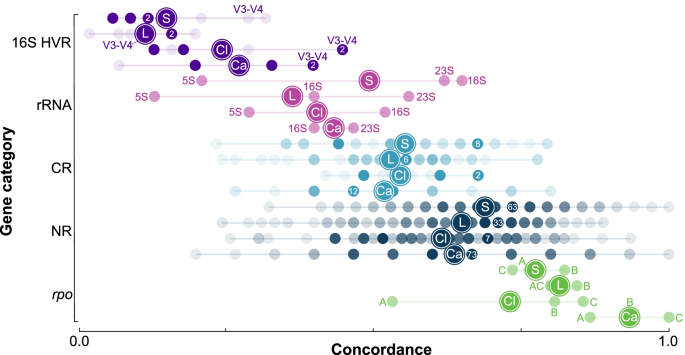
Determining the most accurate 16S rRNA hypervariable region for taxonomic identification from respiratory samples | Scientific Reports

Full-Length 16S rRNA Gene Analysis Using Long-Read Nanopore Sequencing for Rapid Identification of Bacteria from Clinical Specimens | SpringerLink

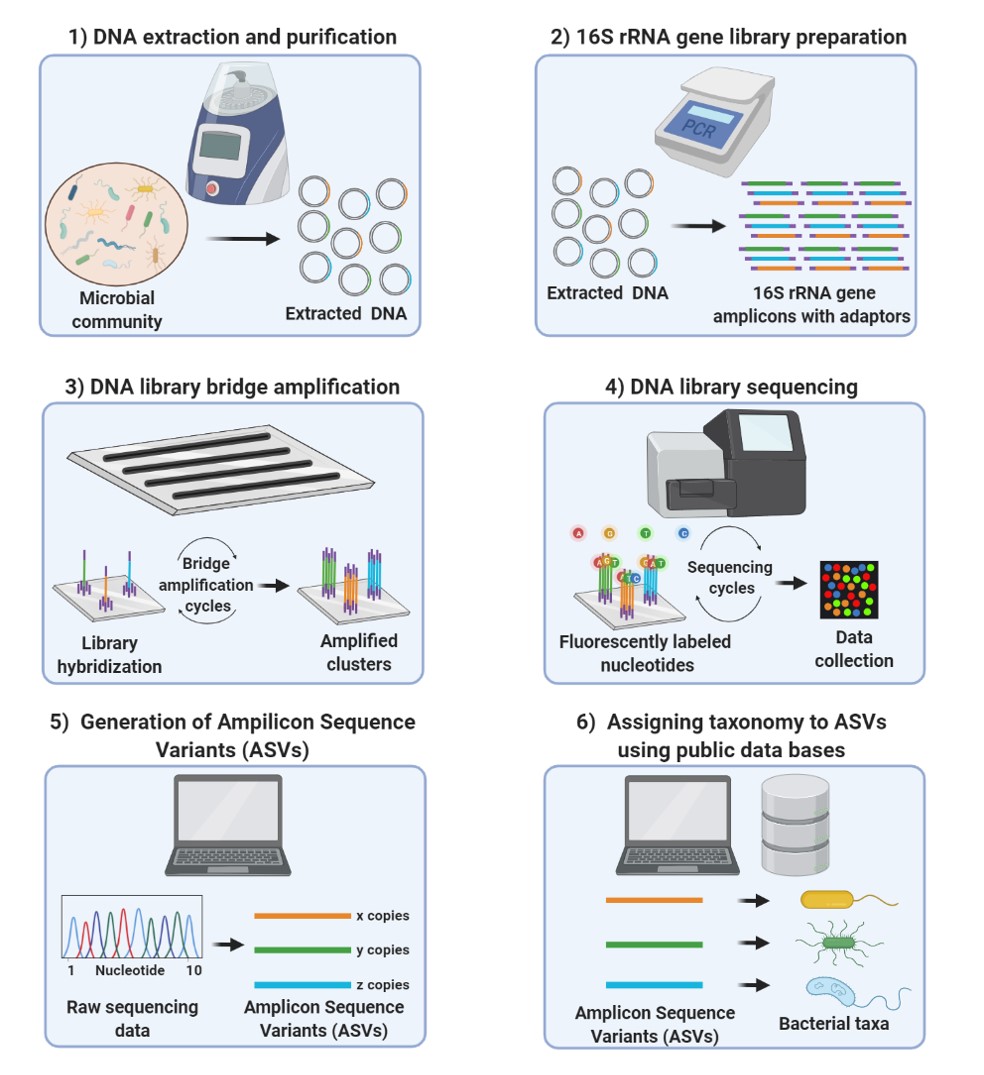
Efficient Nucleic Acid Extraction and 16S rRNA Gene Sequencing for Bacterial Community Characterization | Protocol
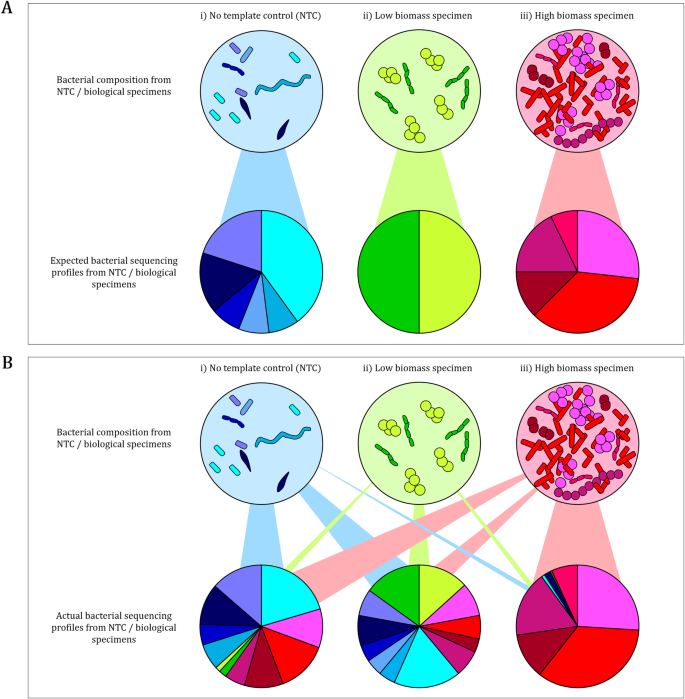
Optimizing 16S rRNA gene profile analysis from low biomass nasopharyngeal and induced sputum specimens | BMC Microbiology | Full Text
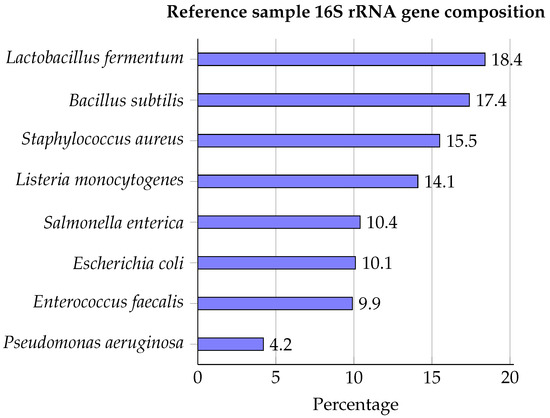
IJMS | Free Full-Text | Targeting the 16S rRNA Gene for Bacterial Identification in Complex Mixed Samples: Comparative Evaluation of Second (Illumina) and Third (Oxford Nanopore Technologies) Generation Sequencing Technologies

16S rRNA gene based analysis of the microbial diversity and hydrogen production in three mixed anaerobic cultures - ScienceDirect
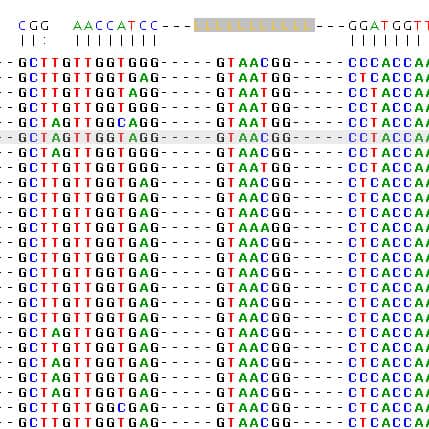
How to calculate 16S rRNA sequence similarity values for bacterial taxonomy: Why BLAST should be avoided – EzBioCloud Help center
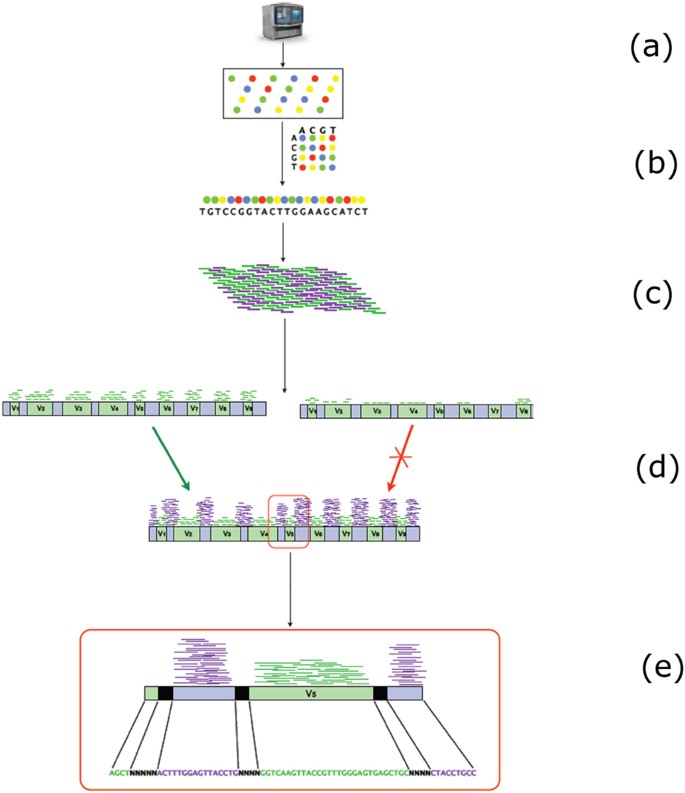
Direct 16S rRNA-seq from bacterial communities: a PCR-independent approach to simultaneously assess microbial diversity and functional activity potential of each taxon | Scientific Reports
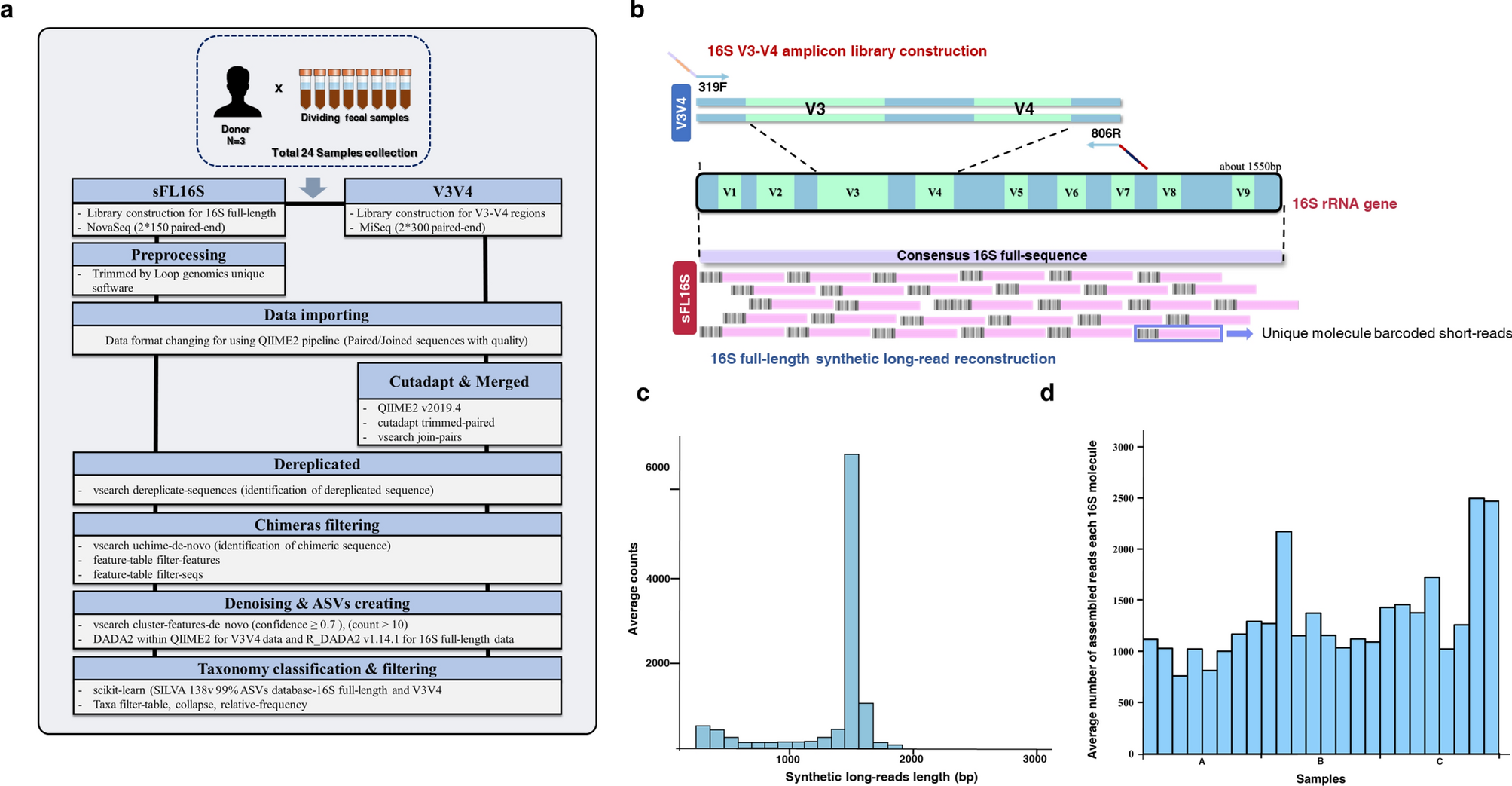
The effect of taxonomic classification by full-length 16S rRNA sequencing with a synthetic long-read technology | Scientific Reports
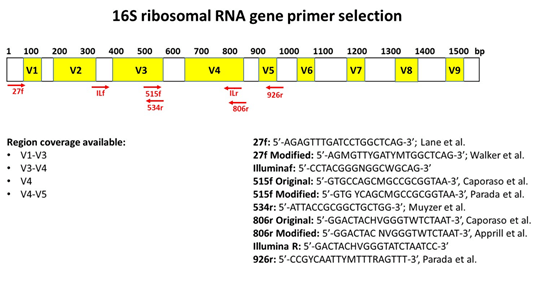
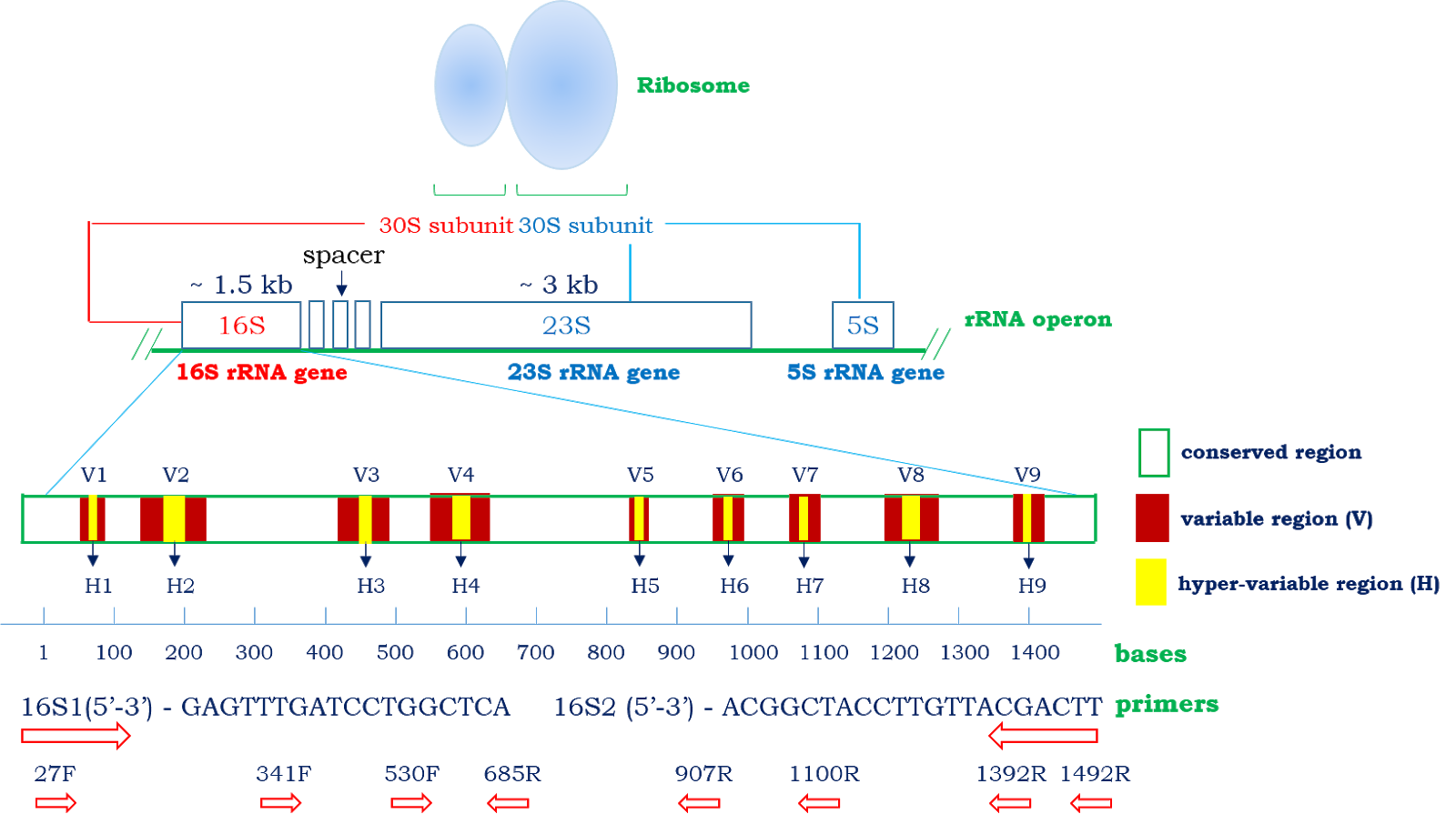
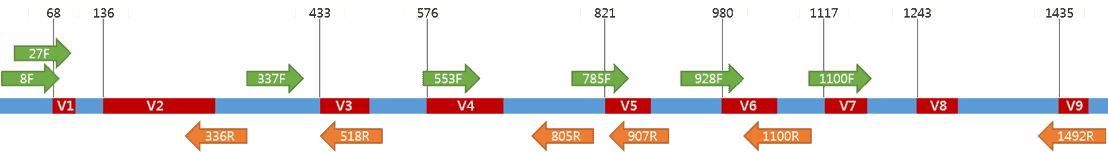
2-16S rRNA gene structure and possible primers A schematic figure of... | Download Scientific Diagram

Full‐length 16S rRNA gene classification of Atlantic salmon bacteria and effects of using different 16S variable regions on community structure analysis - Klemetsen - 2019 - MicrobiologyOpen - Wiley Online Library